16 comments
Awesome summary, thank you Achim.
It would be nice to tie off the story and find out what was the outcome for the question stated in the beginning “look for public available data, describing the social structure of the environment of these schools in question”
Perhaps Achim (or his daughter 🙂 will be willing to share that analysis when they complete it!
Happy you liked it, to compile, analyze and publish the data will be the task of my daughter ;^)
have fun, Achim
Thanks a lot for sharing!!
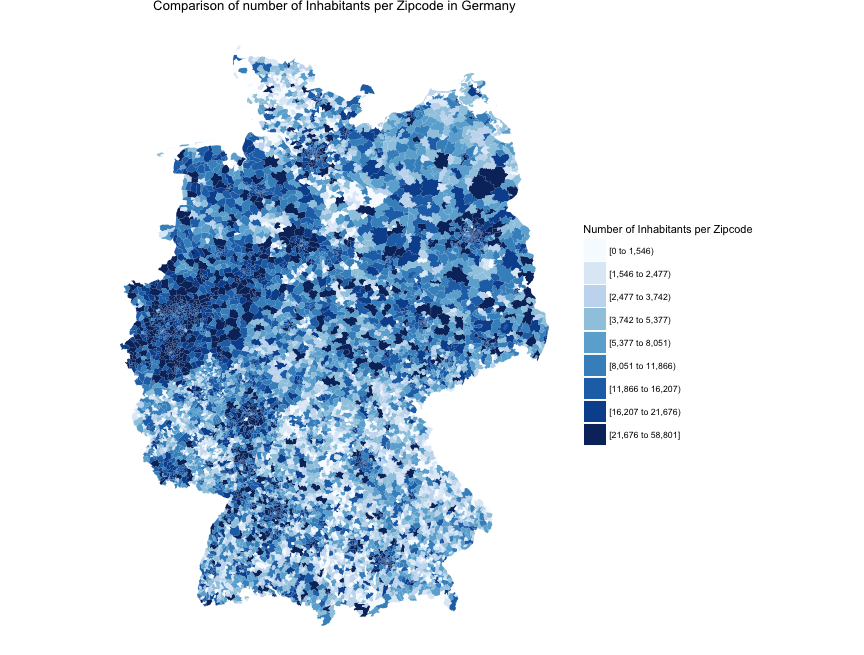
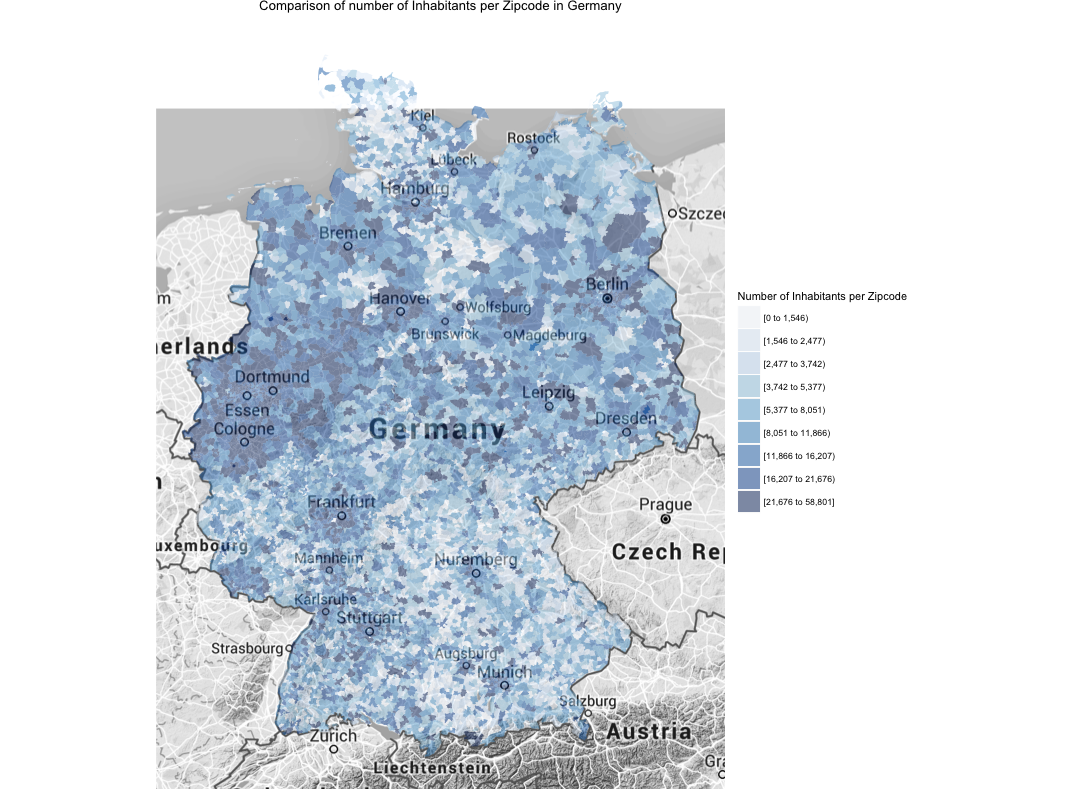
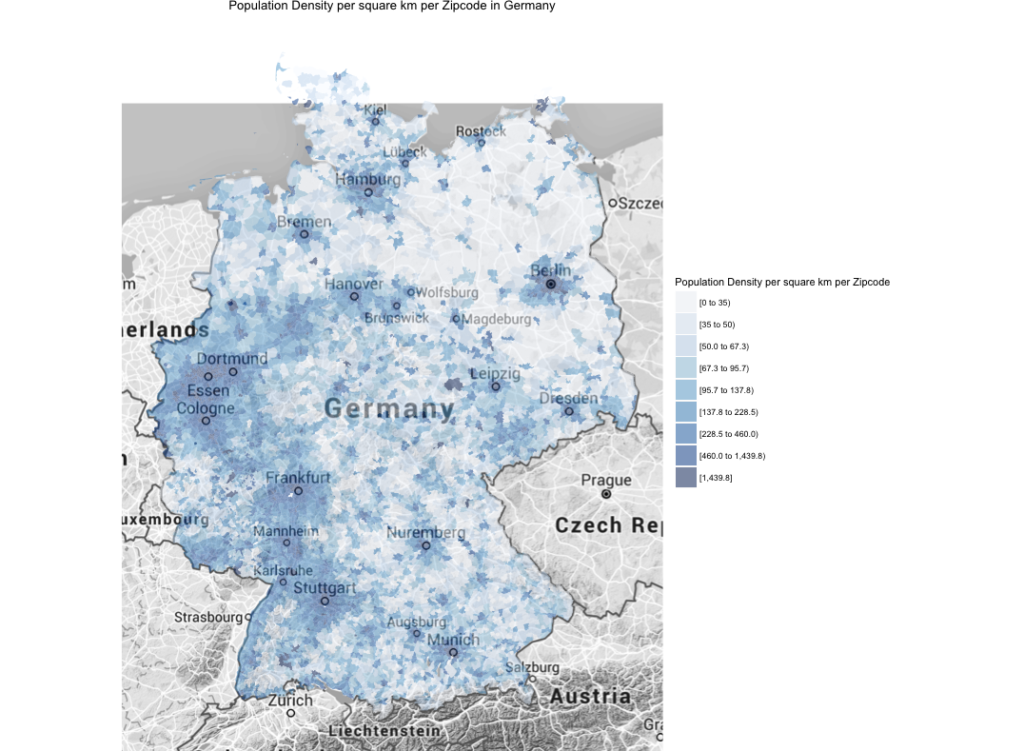
If I understand correctly one can see the number of inhabitants per zip code. Is there a way to also display the density of people per zip code? So the number of inhabitants per m2 of zip code. This would look different since the zip code area size seems to vary a lot wouldn’t it?
I’m also excited about your daughters analysis!
Cheers, Ben
I don’t know about that shapefile, but the ones from the US Census Bureau also have an “area” column, so it is possible there. See this discussion in the choroplethr google group: https://groups.google.com/forum/#!searchin/choroplethr/density/choroplethr/Z6PmsN7xBiM/6Le7nWXiAwAJ
Hi Ben.
Glad you liked my post. You are correct, normalizing the data to inhabitants per m^2 would change the appearance of the map. There is an “area” column buried inside the shapefile structure, I would have to coerce this to change the plot according to your suggestion.
I also looking forward to my daughter analyses, which may take a while, especially acquiring the data.
Have fun,
Achim
Achim – I’ve just come across NUTS 2 and 3 level charts available from Eurostat in another blog post
Here it is – take a look at the charts, very interesting picture emerging about what sounds like may be tangentially related to your daughter’s original line of inquiry. Might be useful to learn about all the data available to capture various dimensions of the impact of the social structure of the environment that you referred to.
Thank you GMS – this looks awesome. Generally it seems easier to get this kind of data from EU sources, than from the German “Statische Landesämter”.
Thanks again, Achim
Awesome tutorial, thanks a lot. When trying to run your code, I receive an error when instantiating the new class saying that “region” is not a map.df colname. I’m a relative beginner and am not sure where exactly the breaking point is. Has anyone else received this error or knows how to solve it?
Hi John.
Glad you liked the post. Sorry you have problems with the example. Do you see this error with the first part of the example, or in the update?
Have fun, Achim
Hi Achim,
thanks a lot for getting back on this. I see this error when trying to execute the very first part of the example when line 47 “c <- GERPLZChoropleth$new(df)" should be executed.
Thanks and best, John
Hi John,
perhaps something went wrong here:
ger_plz.df$region <- ger_plz.df$plz
this command:
head(ger_plz.df)
should give you something like this:
[code lang="r"]
long lat order hole piece id group plz note region
1 5.866315 51.05110 1 FALSE 1 0 0.1 52538 52538 Gangelt, Selfkant 52538
2 5.866917 51.05124 2 FALSE 1 0 0.1 52538 52538 Gangelt, Selfkant 52538
[/code]
have fun
Achim
Thanks, Achim (and Ari by extension) for the great tutorial! One more point to note, R likes to throw the following error when trying to join ‘region x region’: Error: Can’t join on ‘region’ x ‘region’ because of incompatible types (integer / factor). This can easily be solved by making sure both region columns in df and ger_plz.df are either integer/factor (as long as they are the same) I solved it with the as.factor() function.
All best,
Matthias
late to the party, but just wanted to say, these are great. thanks
Hi i am a geographer and R spatial coder from Rio de Janeiro..great post very useful code thks!
Hi, i am very new to this so apologies in advance if the question sounds naive. I copied the script in an editor and ran it using the source command in the R console. Everything seems to work fine except i cannot see the map in quartz, the message i am getting after running it is:
OGR data source with driver: ESRI Shapefile
Source: “.”, layer: “plz-gebiete”
with 8712 features
It has 2 fields
Parsed with column specification:
cols(
plz = col_character(),
einwohner = col_integer()
)
Map from URL : http://maps.googleapis.com/maps/api/staticmap?center=50.233703,10.362028&zoom=6&size=640×640&scale=2&maptype=terrain&language=en-EN&sensor=false
Scale for ‘x’ is already present. Adding another scale for ‘x’, which will replace the existing
scale.
Scale for ‘y’ is already present. Adding another scale for ‘y’, which will replace the existing
scale.
Warning messages:
1: In gpclibPermit() :
support for gpclib will be withdrawn from maptools at the next major release
2: In left_join_impl(x, y, by$x, by$y, suffix$x, suffix$y) :
joining character vector and factor, coercing into character vector
3: In left_join_impl(x, y, by$x, by$y, suffix$x, suffix$y) :
joining character vector and factor, coercing into character vector
Any idea why?
Thanks for taking the time to post this code and detail all the steps.